Top 10 Progress in China's Life Sciences in 2022 released
On January 19, the Consortium of Life science Societies of the China Association for Science and Technology announced the top ten advances in China's life sciences in 2022. Ten research advances in the field of life sciences, including the research on "immune escape mechanism of novel coronavirus mutant strains" carried out by the Cao Yunlong Institute of Peking University, were included in the list.
According to reports, the ten advances include 7 knowledge innovation and 3 technological innovation project results, which are recommended by the member societies of the organization, selected by senior experts in the fields of basic life science, biotechnology and clinical medicine, and finally determined after the review of the Presidium of the Life Science Association of the China Association for Science and Technology.
According to the data provided by the China Association for Science and Technology, most of the results of the ten progressing projects have been published in the three top international journals CNS (Cell, Nature, Science), reflecting the outstanding originality of China's life science research results.
In addition, many of the results are multi-level and multi-angle disassembly and exploration of the same scientific problem in the way of paper serialization, showing the system and depth of life science research.
For example, the study on the immune escape mechanism of COVID-19 mutant strains analyzed the structural and infectious characteristics of multiple mutant strains for the first time, and detailed the full epitope distribution and escape map of COVID-19 neutralizing antibodies, which were divided into five papers. Based on autonomous DNA nanosphere sequencing technology, the team led by Wang Jian and Xu Xun of the BSU Institute of Life Sciences has completed a high-precision panoramic space-time gene expression map of life completed by several research teams in the United Nations, and has published five papers in the form of special topics and cover articles in journals such as Cell and Science.

The achievements in the ten major developments highlight the responsibility and mission of scientific research to shoulder "facing people's life and health". Among them, the study of "ischemic cerebrovascular disease precision treatment program" has created a precedent in the international field of cerebrovascular disease precision treatment; The study of "molecular mechanism of initiation of autophagy in multicellular organisms" is of great significance for exploring the occurrence and development of neurodegenerative diseases.
The relevant person in charge said that the current selection activities have been carried out for eight consecutive years. After the selection results are announced, the selected project experts will be invited to write and publish popular science books, and the new mysteries of life science will be revealed to the public through popular science reports. The ten major advances will also provide new ideas for the development of new life science technologies, new medical breakthroughs and the development of bioeconomy.
China's top 10 Progress in life Sciences in 2022
1. Immune escape mechanism of novel coronavirus mutant strains
The Omicron strain of the new coronavirus continues to mutate, causing multiple rounds of outbreaks around the world. Resolving the humoral immune escape mechanism of COVID-19 mutants is of great guiding significance for the development of COVID-19 vaccines and epidemic prevention and control.
Xiaoliang Xie and Yunlong Cao from Peking University, together with Xiangxi Wang from the Institute of Biophysics of the Chinese Academy of Sciences and Youchun Wang from the Chinese Institute of Food and Drug Control, were the first to report the humoral immune escape characteristics and molecular mechanisms of COVID-19 omicrone and its subtype variants. For the first time, the structural and infective characteristics of several mutant strains were analyzed, and the full epitope distribution and escape map of COVID-19 neutralizing antibodies were described in detail. It was revealed that the mutation carried by Omicron BA.1 can specifically escape the neutralizing antibodies induced by the original strain infection and vaccination, while the mutation carried by Omicron BA.4/BA.5 can specifically escape the neutralizing antibodies induced by BA.1 infection, proving that herd immunity through Omicron infection can not be achieved to block the transmission of COVID-19. These studies have enhanced the scientific understanding of COVID-19 prevention and control in the world, and provided important data reference and theoretical support for the research and development direction of broad-spectrum COVID-19 vaccines and antibody drugs.
2. New pathways of cholesterol efflux and new strategies for lipid reduction
There are 330 million patients with cardiovascular disease in China, and excessive cholesterol in the blood is the main risk factor. Although existing lipid-lowering drugs can reduce lipids to different degrees, they have certain side effects and limitations. The molecular structure of cholesterol determines that it is difficult to be degraded in the body, so the discovery of the method of expelling cholesterol to the body is of great significance for the development of new lipid-lowering drugs.
Song Baoliang's team at Wuhan University Taikang Life Medical Center found that after the loss of the glycoprotein receptor ASGR1, cholesterol is excreted into the bile and further left the body through the stool. Inhibition of ASGR1 function can promote a large number of cholesterol effluents, blood lipids and liver fat decrease, and play a good effect on atherosclerosis. At the same time, the neutralizing antibody of ASGR1 can be combined with existing lipid-lowering drugs to play a better lipid-lowering effect. This finding points the way for the development of new lipid-lowering drugs to promote cholesterol effluents, and ASGR1 has become a hot target for many pharmaceutical companies to develop lipid-lowering drugs.
Inhibition of ASGR1 promotes cholesterol excretion into bile and stool and prevents atherosclerotic plaque formation Inhibition of ASGR1 promotes cholesterol excretion into bile and stool and prevents atherosclerotic plaque formation
3. New technologies of mammalian chromosome engineering and artificial evolution of chromosomes
During the long evolution of life, chromosomal rearrangement occurs, resulting in karyotype variation. Rodents accumulate 3.2-3.5 chromosome rearrangements every million years, while primates accumulate 1.6. How can such events be simulated and studied in laboratory model animals?
The team of Li Wei and Zhou Qi from the Institute of Zoology of the Chinese Academy of Sciences, together with the team of Li Jinsong from the Innovation Center for Molecular and Cell Science of the Chinese Academy of Sciences, achieved the programmable connection of complete chromosomes of mammals for the first time, creating a series of experimental mice with a new karyotype of 19 pairs of chromosomes. Events of karyotype evolution that take hundreds to tens of thousands of years to achieve in nature have been achieved in the laboratory by artificial design. The study found limitations in chromosome length; The effect of chromosome rearrangement on reproduction was revealed. It is confirmed that the robustness of genome assembly is an important basis for chromosome evolution, which provides a feasible technical route for the modification of mammalian chromosome structure, the creation of new karyotype subspecies and the simulation of diseases with chromosome structural variation, and opens a new field of mammalian chromosome genetic modification.
Chromosome connected mouse "Xiao Zhu" has a unique chromosome type Chromosome connected mouse "Xiao Zhu" has a unique chromosome type
4. Study on translation group map of early human embryo and activation factors of zygotic genome
After fertilization of the human egg, the early embryo is basically in a state of transcriptional silence at first, and translation regulation plays an important role in ovum maturation, fertilization and embryo genome activation. As the first gene expression of life, zygotic genome activation is a landmark event in the initiation of embryonic development. However, how the human zygotic genome is activated has long been an unsolved mystery.
The research team of Professor Wei Jie of Tsinghua University, Academician Zijiang Chen and Professor Han Zhao of Shandong University has mapped the translation map of early human embryo development for the first time by developing the combined sequencing technology of ultra-sensitive translation group and transcriptome. This work identified TPRX1/2/L family proteins by searching for transcription factors with high translation during genome activation, demonstrating that they play an important regulatory role in human zygotic genome activation and early embryonic development. The work addresses major fundamental scientific questions about how human embryo procedures were first initiated, and provides an important theoretical foundation and research tool for future treatments of infertility and improved assisted reproductive technologies.
The human egg and early embryo translation group and transcriptome were jointly sequenced to reveal the translation regulation mechanism and key factors of zygotic genome activation
5. High-precision panoramic space-time gene expression mapping of life
Cells are the basic functional units of life. Analysis of cell type, localization, and intercellular communication is critical to understanding organ function, ontogeny, human disease, and organ evolution in species.
Based on autonomous DNA nanosphere sequencing technology, a team led by Wang Jian and Xu Xun from BGI Institute of Life Sciences developed high-precision large-field spatial transcriptome technology, which promoted the resolution of understanding life to the sub-cellular level of 500nm, which increased the resolution by 200 times and the size of the field of view by 483 times compared with similar technologies in the past. Based on this technology, a team from BTU, the Chinese Academy of Sciences, Southern University of Science and Technology, Huazhong Agricultural University and Guangdong Provincial People's Hospital has mapped for the first time in the world the highest precision and most comprehensive spatio-temporal gene expression dataset to date for important model organisms such as mice, fruit flies, zebrafish, Arabidopsis and salamander, and discovered new cell types that play a key regulatory role in the process. After the publication of this series of results, the international community has aroused a warm response, promoting the establishment of STOC, a global alliance of spatiotemporal omics led by Chinese scientists, which has attracted the participation of more than 190 research teams from 25 countries.
6. Discovery of metformin targets and elucidation of its mechanism of delaying aging
Metformin is not only the first-line drug for the treatment of type 2 diabetes, clinical studies have also found that metformin has anti-tumor, anti-aging and other magical effects. But metformin has been on the market for 65 years and its target has remained a mystery.

Xiamen University Lin Shengcai team after 7 years of scientific research, found a protein called PEN2 is the target protein of metformin. Importantly, this study not only identified the direct target of metformin, but also mapped out the road map of metformin's function from a molecular perspective. They also screened a chemical drug (commonly known as "Gu Jing") that can simulate the effect of Gu Gu (calorie restriction), which has the effect of lowering sugar, treating fatty liver, and extending life span. It was also found that "talisman" and metformin both use the glucose (calorie restriction) sensing pathway previously discovered by them, so as to couple to AMPK longevity related pathway, to treat major metabolic diseases such as diabetes and fatty liver and delay aging. Kei Sakamoto, an expert on metformin's mechanism of action, and Niels Jessen, a diabetes researcher, wrote that the discovery has important value in developing new alternative drugs to overcome metformin's defects. AMPK field expert David Carling has written that "talisman" is the magic bullet in the long-sought fight against metabolic diseases.
7. Precise treatment of ischemic cerebrovascular disease
High recurrence is a global problem in the prevention and treatment of ischemic cerebrovascular diseases. However, the effect of aspirin single antiplatelet therapy recommended by national guidelines is limited, and clinical studies of combined antiplatelet therapy with other drugs have failed due to ineffectiveness or increased risk of serious bleeding. Therefore, combined therapy has been banned for ischemic cerebrovascular diseases by international guidelines.
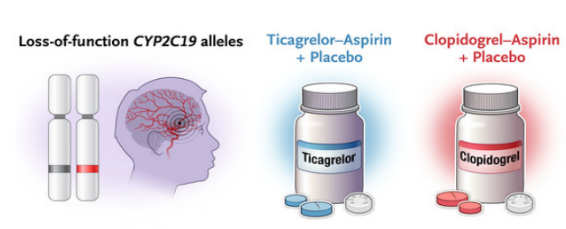
Wang Yongjun's team of Beijing Tiantan Hospital affiliated to Capital Medical University proposed for the first time in the world the short-range double-channel double-effect combination treatment program of aspirin plus clopidogrel, which rewrote the guidelines of many countries such as Europe and the United States. Based on this regimen, he found that ABCB1, CYP2C19 and F2R, the key genes of clopidogrel absorption and metabolic pathways, all significantly affect the efficacy of the drug, and proposed a "bypass gene" ticagrelor replacement therapy regimen for people carrying loss-function alleles of clopidogrel, which can reduce the risk of recurrence by 23%. It was evaluated by NEJM as opening a new era of gene-guided treatment of cerebrovascular diseases, and was evaluated by Nature Medicine and European Heart Journal as a new opportunity for genotyping to guide the treatment of cerebrovascular diseases.
The findings were published in the New England Journal of Medicine.
Efficacy of ticagrelor replacement therapy in persons carrying CYP2C19 function loss alleles
8. Developed subversive gene decoding technology to describe the world's first "disturbance map"
The human genome has long been sequenced, but its function is still poorly understood, which seriously hinders the diagnosis and treatment of diseases. The Chi Tian team of Shanghai University of Science and Technology has developed a unique path and fused the two underlying tools of "CRISPR gene editing" and "Cre gene recombination" into a subversive "high-throughput, pan-organization" gene function decoding technology iMAP, which can increase the decoding speed of mouse genes by at least 100 times. This work also uses iMAP to successfully describe the world's first "disturbance map", which shows the basic functions of 90 protein-coding genes in 39 kinds of tissue cells in mice, and will inevitably give birth to a "panoramic disturbance map" covering all genes and tissues and decoding the entire "book of life", which will become an indispensable "world map" for people to explore the mysteries of life in the future. iMAP is robust, easy to operate, easy to popularize, and has a wide range of uses (including mining disease drug targets and optimizing rice varieties), and has achieved technological breakthroughs in the field of gene decoding from 0 to 1.
How iMAP works and its uses. The image on the right (" Flying Flowers ") is a "Dunhuang version" of how iMAP works, showing the flying sky (Cre) releasing flowers (gRNA) from a vase (genetically modified), which produces chimeric mice (design: Li Zhuoshi, Chi Tian).

9. Molecular mechanism of initiation of autophagy in multicellular organisms
Autophagy acts as a "scavenger" in cells by wrapping misfolded proteins, damaged organelles and other "garbage" in a double-membrane structure called the autophagosome and transporting it to the lysosome for degradation and recycling. Autophagy is essential for resistance to many kinds of stress and maintenance of cell homeostasis. Finding the signal that determines the formation of autophagosomes is a long-standing problem in the field of autophagy.
Zhang Hong and his team from the Institute of Biophysics of the Chinese Academy of Sciences found that during autophagy induction, calcium transients occur on the surface of the endoplasmic reticulum, triggering liquid-liquid separation of the FIP200 autophagy initiation complex, and the formed FIP200 condensate binds to the ER protein and is located in the endoplasmic reticulum, becoming the initiation site of the autophagy. This work has revealed that calcium transient on the surface of the ER is a key signal to initiate the formation of autophagosomes, greatly promoting the understanding of the molecular mechanism of autophagy, and has important significance for exploring the mechanism of autophagy abnormalities in neurodegenerative diseases caused by calcium disorders in the ER.
Calcium transients on the surface of the endoplasmic reticulum are a key signal for autophagosome formation
10. New mechanism of gene mining and regulation of high temperature resistance in rice
With the global warming, extreme hot weather occurs frequently, which greatly reduces crop production and aggravates food security problems. It is an urgent challenge to explore the gene resources of crop resistance to high temperature and elucidate the regulation mechanism of high temperature resistance so as to breed crop varieties resistant to high temperature.
The team led by Hongxuan Lin of the Center of Excellence in Molecular Plant Science at the Chinese Academy of Sciences (CAS) and Youshun Lin of Shanghai Jiao Tong University (SJTU) has revealed a new mechanism of high temperature resistance in rice, unearthed a genetic module TT3 composed of TT3.1 and TT3.2, and discovered the first potential high temperature receptor (TT3.1) for the first time. It senses and transmits high temperature signals to the chloroplast protein TT3.2, which protects the chloroplast from heat damage.
- ABB
- General Electric
- EMERSON
- Honeywell
- HIMA
- ALSTOM
- Rolls-Royce
- MOTOROLA
- Rockwell
- Siemens
- Woodward
- YOKOGAWA
- FOXBORO
- KOLLMORGEN
- MOOG
- KB
- YAMAHA
- BENDER
- TEKTRONIX
- Westinghouse
- AMAT
- AB
- XYCOM
- Yaskawa
- B&R
- Schneider
- Kongsberg
- NI
- WATLOW
- ProSoft
- SEW
- ADVANCED
- Reliance
- TRICONEX
- METSO
- MAN
- Advantest
- STUDER
- KONGSBERG
- DANAHER MOTION
- Bently
- Galil
- EATON
- MOLEX
- DEIF
- B&W
- ZYGO
- Aerotech
- DANFOSS
- Beijer
- Moxa
- Rexroth
- Johnson
- WAGO
- TOSHIBA
- BMCM
- SMC
- HITACHI
- HIRSCHMANN
- Application field
- XP POWER
- CTI
- TRICON
- STOBER
- Thinklogical
- Horner Automation
- Meggitt
- Fanuc
- Baldor
- SHINKAWA
- Other Brands
- UniOP
- KUKA
- Iba






































































































































